Sarah Lott Moretto
Conference 2022 Poster Presentation
Project title
In-house RT-PCR application for detection of SARS-CoV-2.
Authors and Affiliations
Sarah Lott Moretto1, Maria Angelica Ehara Watanabe1, Mateus Nobrega Aoki2, Marla Karine Amarante1, Karen Brajão de Oliveira1, Carolina Batista Ariza3,4
1. Department of Pathological Sciences, State University of Londrina, Londrina, Brazil.
2. Laboratory of Applied Sciences and Technologies in Health, Carlos Chagas Institute, FIOCRUZ, Curitiba, Brazil.
3. Department of General Biology, State University of Londrina, Londrina, Brazil.
4. Filadelfia University Center, Biomedicine Collegiate, Londrina, Brazil.
Abstract
Background
The disease caused by the SARS-CoV-2 virus can present mild to severe symptoms. COVID-19 has been proving to be a challenging disease for moderne medicine. With an ideal treatment not yet established and because of its potential for mortality both in individuals with and without comorbidities, the detection of the virus and diagnosis of the disease made as soon as possible can help in a more targeted and effective treatment, contributing to a faster recovery of the patient. Samples from critically ill patients suspected of COVID-19 are being tested to have the correct diagnosis of the disease. The work, which was initially being carried out only in the Central Public Health Laboratories and Public Reference Laboratories, is now being gradually expanded through the efforts of the brazilian Ministry of Health. The disease has been detected through the detection test of SARS-CoV-2 by real time PCR (Polymerase Chain Reaction in real time) which requires medium-sized equipment that can be found in laboratories covering the area of molecular biology. The purpose of this work was to evaluate primers from specific regions of the virus and perform conventional RT-PCR, a fast, low-cost method which can be performed with few samples in any laboratory routine in any region of Brazil.
Methods
This study was approved by the Ethics Committee for Research involving human beings at the State University of Londrina (CAAE 36247920.1.0000.5231). In the present work, viral RNA from 106 copies/ul culture and primers for different regions of SARS-CoV-2 Spike (S), nucleocapsid (N), membrane glycoprotein (M) and RNA-dependent RNA polymerase (RdRp) of SARS-CoV-2 – ID: C2H0GV2V014 of the viral genome were used. Real-time PCR was performed with these primers through reverse transcription reaction and PCR with GoTaq qPCR and conventional PCR, in which the amplicons were visualized by polyacrylamide gel electrophoresis, stained with silver nitrate.
Results
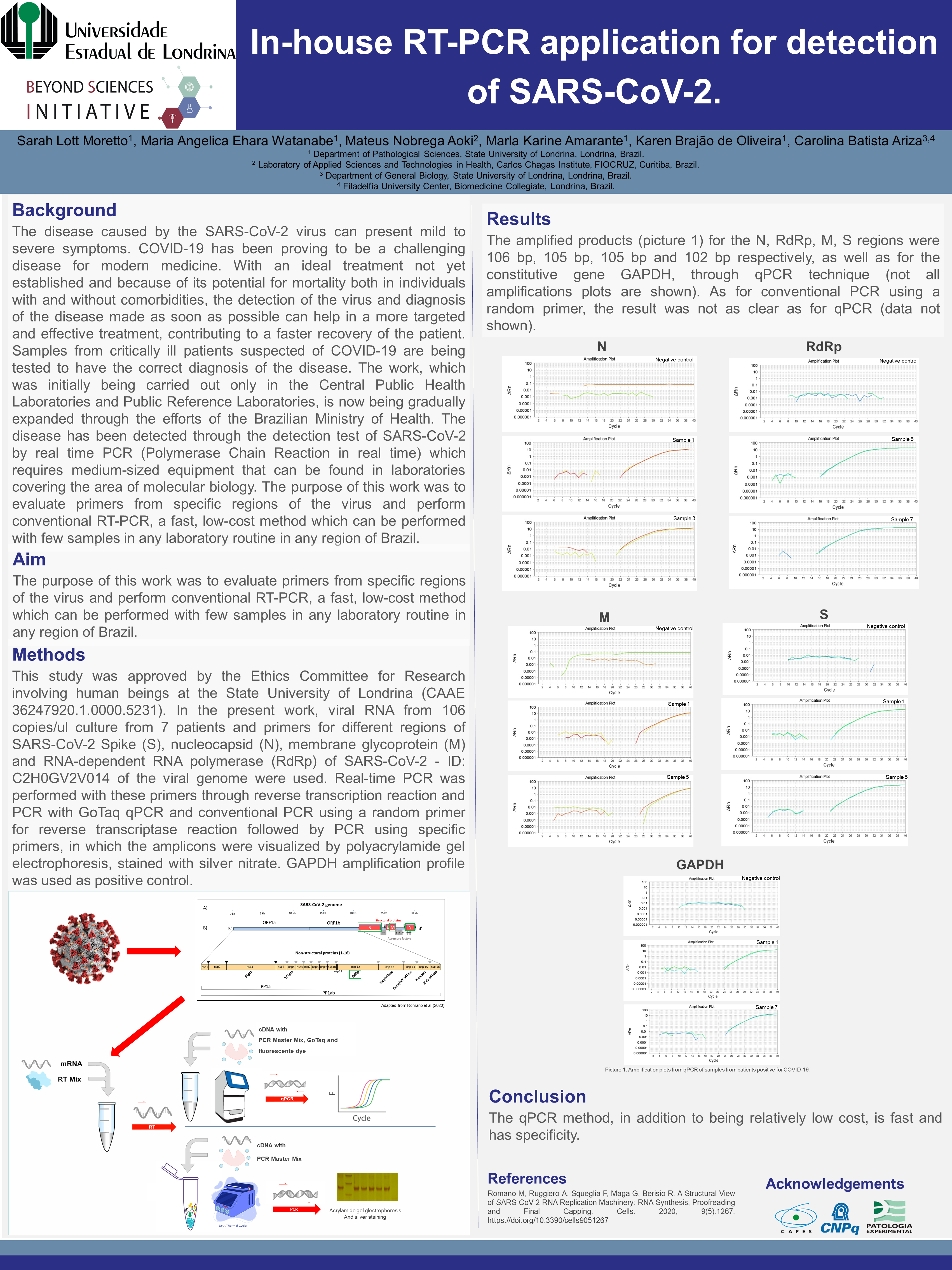
The amplified products for the S, N, M and RdRp regions were 106 bp, 105 bp, 105 bp and 102 bp respectively.
Conclusions
This method, in addition to being low cost, is fast and has specificity.