Tanya Jindal
India
“In Silico Discovery of phytochemicals Inhibiting Bacterial Urocanate Reductase to Prevent Imidazole Propionate-Mediated Insulin Resistance in T2D”
Tanya Jindal1, Rohit Bansal1, Preeti Kumari1, Archana Bhatnagar1
1. Department of Biochemistry, Panjab University, Chandigarh, India.
Abstract
Background
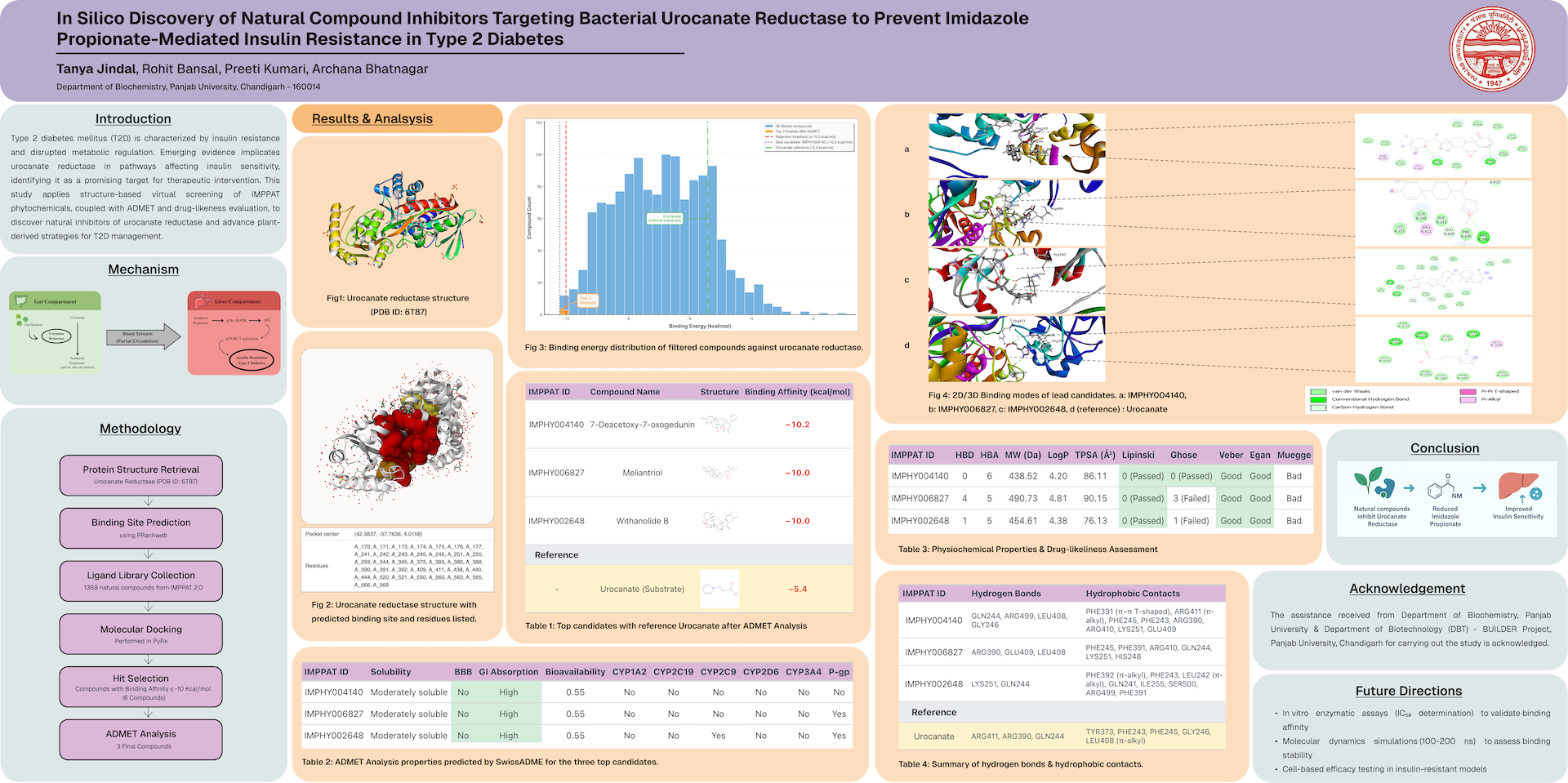
Type 2 Diabetes Mellitus (T2DM) is a complex metabolic disorder primarily associated with insulin resistance and impaired glucose metabolism. Recent studies implicate gut microbiota–derived imidazole propionate as a metabolite that disrupts insulin signaling. The bacterial enzyme urocanate reductase (UrdA) catalyzes the formation of imidazole propionate and has thus emerged as a potential therapeutic target. Targeting UrdA with natural inhibitors may help mitigate microbial contributions to insulin resistance in T2DM.
Methods
The crystal structure of UrdA (PDB ID: 6T87) was retrieved and analyzed to predict active binding sites using PrankWeb. A phytochemical library from IMPPAT 2.0 comprising 1,359 natural compounds was subjected to virtual screening and molecular docking using PyRx. Compounds with binding energies ≤ –10 kcal/mol were shortlisted. The top hits were further evaluated for ADMET properties using SwissADME to assess pharmacokinetic profiles and drug-likeness.
Results
Three lead compounds—IMPHY004140 (7-Deacetoxy-7-oxogedunin), IMPHY006827 (Meliantriol), and IMPHY002648 (Withanolide B)—showed high binding affinities ranging from –10.0 to –10.2 kcal/mol, outperforming the natural substrate urocanate (–5.4 kcal/mol). These ligands exhibited key hydrogen bonding and hydrophobic interactions with catalytic residues ARG391, GLN244, and PHE243. ADMET analysis predicted good solubility, high gastrointestinal absorption, and acceptable bioavailability, confirming their drug-like characteristics.
Conclusions
The in silico identification of potent natural inhibitors of urocanate reductase highlights their potential to reduce imidazole propionate production and improve insulin sensitivity. This computational approach provides a foundation for future in vitro enzyme inhibition assays and molecular dynamics simulations to validate stability and efficacy in T2DM therapy.

Leave A Comment